FAQ about microscopy
Microscopy questions
For a good widefield deconvolution it is not necessary to acquire large regions above and below the in-focus zone. In widefield microscopy the blur can spread through large regions. The Huygens algorithms take this into account, thus imaging very far above and below the in-focus region is not necessary to achieve good deconvolution results. The image can be cropped far from the in-focus region to speed up the deconvolution process. To avoid cropping slices that contain useful information, leave a border of around 1-2 Half Intensity Width sizes. As an example, in widefield, well-sampled images recorded with NA 1.3 this corresponds to 0.75 to 1.5 microns.
Piezo driven focussing systems are also not without problems: hysteresis, temperature drift to name a few. To compensate for the latter, especially in cases where experiments are to be conducted at 37 degrees, we recommend a sensor feedback system.
If on the other hand you are using an RGB camera, the situation is a bit different. Each of the camera components has its own, probably not so sharp band filter. The camera filters are in cascade with the microscope bandfilter. Suppose the microscope filter passes all wavelengths longer than 500nm, and the blue camera filter passes all shorter than 500nm, then the Blue channel will be dark. In a different case, suppose the microscope filter passes all wavelength longer than 570nm, the green camera filter from 500-600nm, and the red camera filter from 580nm and longer. Then both Green and Red channel contain an image. With little loss of accuracy these could be added and deconvolved at 570nm emission wavelength (you need Huygens Pro for that).
$$r_{nm} = \frac{d_{mm} \cdot 10^6}{2 \cdot M_{sys} \cdot M_{obj}}$$ with r the backprojected radius in nm, d the pinhole diamater in mm, M_{sys}$$ the system magnification, M_{obj}$$ the objective magnification. The "2" is for diameter to radius conversion. Depending on the configuration the system Magnification for the MRC 600 or 1024 is 53-83. The system magnification of the Radiance is reported to be 60.
For more information and a calculator you can visit BackprojectedPinholeCalculatorIf you specify an NA which is higher than the r.i. of the medium, then this implies total internal reflection at the glass/medium interface, causing the oblique angles of the excitation light to be reflected back into the microscope, thereby reducing the effective NA.
See also [TotalInternalReflection|TotalInternalReflection].Imaging diatoms is indeed a difficult task. This is so because the skeleton of diatoms has a high refractive index and can seriously spoil light propagation, resulting in a PSF which varies strongly from place to place.
Yes! If you enter a figure not much lower than the NA you'll see it gets highlighted to warn you against an optically poor situation.
A factor which can increase this elongation considerably is refractive index mismatch of embedding medium and coverslip. If this is the case and you can't increase the refractive index of the medium you might consider purchasing a water or glycerin immersion objective. To get a more accurate estimate of what the bead should look like in the microscope, you can simulate the imaging process with Huygens Professional:
- start Huygens Pro
- generate a sphere + do an FFT on the result
- generate a PSF + do an FFT on the result
- multiply the two and put the result in say image 'c'
- do a reverse FFT on 'c' (with the FFT tool in auto mode this will go automatically)
- do a 'to/from optical representation' (restoration menu) to shift the origin of the image to the center.
- to make things realistic you can throw in some Poisson noise.
The pinhole measure as used by Huygens is the "backprojected pinhole RADIUS", which means that the physical size should by divided by the systems magnification factor. Using a 100x objective lens the value is 410 nm, which is still pretty high (250 nm is a common value), so maybe there is some more internal magnification in the system we are not aware of.
backprojected_RADIUS = number_of_Airy_disks * 0.61 * $$\frac{\lambda}{NA}$$ NA = the numerical aperture, number_of_Airy_disks = the number of disk diameters fitting in the pinhole diameter.
The Airy disk diameter is the diameter of the first dark ring of the diffraction pattern in the focus of a lens, the so-called Airy disk.Example: with an NA = 1.4 and a watery medium (refr. index slightly above 1.3), the most oblique rays cannot pass into the medium and will be totally reflected, effectively limiting the NA of the objective.
See also: What does the message: 'total reflection at glass/medium interface (...)' mean?
If there is a non-descanned detector (NDD), or a very large (say > 10 Airy disks) pinhole, the contribution of the pinhole to the image formation can be ignored. In this case the microscope type can be widefield in combination with 2p photon. This will result in more realistic values for the Nyquist rate than a confocal setting.
Fairly large pinhole
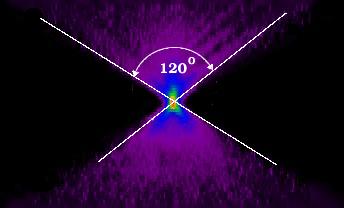
I use gelatin to simply immobilize my sample (suspend it in place in 3-dimensions) which will otherwise tend to "swim away" in an aqueous medium. I do not notice any autofluorescence in the 5% gelatin.
(Answer by Harry Leung, M.Sc.,Technical Officer, Zoology Department, University of Western Ontario, London, Ontario N6A 5B7, E-mail: leung at julian.uwo.ca)In practice, depending on wavelength, for a 1.3 NA objective this corresponds to a backprojected pinhole radius of 200nm to 300nm.
